Department of Chemistry
University of Illinois at Chicago
Chicago IL 60607
Research Overview
Research in the Lorieau group integrates Biophysics,
Physical Chemistry, Structural Biology and Biochemistry in
elucidating the interplay between biomolecular structure,
dynamics, chemistry and function.
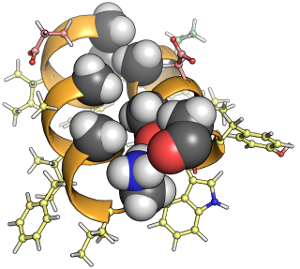
With a combination of solution- and solid-state Nuclear Magnetic Resonance spectroscopies, computational tools and other biophysical methods, our research focuses on membrane protein structure and dynamics, membrane protein biochemistry, the development of theory and techniques to enhance the precision and resolution of structural and dynamic information by NMR, and the investigation of molecular dynamics as it relates to enzymatic catalysis and kinetics.
Membranes and Membrane Proteins
High-Resolution NMR Methods Development
Congratulations to Dr. Charles F. DeLisle!
 Congratulations to Charles F. DeLisle on earning his PhD! Dr. DeLisle
solved the first structure of human proamylin in micelles using
solution NMR. He has made important contributions in our lab by
solving solution NMR structures of membrane proteins, collecting
high-resolution NMR structural data and refining structures with
RDCs. He will be joining New Equilibrium
Biosciences as a Biophysical
Scientist.
Congratulations to Charles F. DeLisle on earning his PhD! Dr. DeLisle
solved the first structure of human proamylin in micelles using
solution NMR. He has made important contributions in our lab by
solving solution NMR structures of membrane proteins, collecting
high-resolution NMR structural data and refining structures with
RDCs. He will be joining New Equilibrium
Biosciences as a Biophysical
Scientist.
Pro‐islet Amyloid Polypeptide in Micelles Contains a Helical Prohormone Segment
Pro‐islet amyloid polypeptide (proIAPP) is the prohormone precursor molecule to IAPP, also known as amylin. IAPP is a calcitonin family peptide hormone that is cosecreted with insulin, and largely responsible for hunger satiation and metabolic homeostasis. Amyloid plaques containing mixtures of mature IAPP and misprocessed proIAPP deposit on, and destroy pancreatic β‐cell membranes, and they are recognized as a clinical hallmark of type 2 diabetes mellitus. In order to better understand the interaction with cellular membranes, we solved the solution NMR structure of proIAPP bound to dodecylphosphocholine micelles at pH 4.5. We show that proIAPP is a dynamic molecule with four α‐helices. The first two helices are contained within the mature IAPP sequence, while the second two helices are part of the C‐terminal prohormone segment (Cpro). We mapped the membrane topology of the amphipathic helices by paramagnetic relaxation enhancement, and we used CD and diffusion‐ordered spectroscopy to identify environmental factors that impact proIAPP membrane affinity. We discuss how our structural results relate to prohormone processing based on the varied pH environments and lipid compositions of organelle membranes within the regulated secretory pathway, and the likelihood of Cpro survival for cosecretion with IAPP.
Partial alignment, residual dipolar couplings and molecular symmetry in solution NMR
Residual dipolar couplings (RDCs) and residual anisotropic chemical shifts (RACSs) are produced by the partial alignment of solution NMR samples. RDCs and RACSs yield high-resolution structural and dynamic information on the orientation of bonds and chemical groups in molecules. Many molecules form oligomers or have intrinsic symmetries, which may simplify the analysis of their partial alignment datasets. In this report, we explore the theory of partial alignment using an irreducible spherical representation, and we investigate the impact of molecular symmetry on the alignment of molecules. Though previous studies have reported simplified relationships on the partial alignment of molecules bearing different symmetry groups, we show that these simplified relationships may not be universal and only apply to a limited set of systems.
Super resolution NOESY spectra of proteins
Spectral resolution remains one of the most significant limitations in the NMR study of biomolecules. We present the srNOESY (super resolution nuclear Overhauser effect spectroscopy) experiment, which enhances the resolution of NOESY cross-peaks at the expense of the diagonal peak line-width. We studied two proteins, ubiquitin and the influenza hemagglutinin fusion peptide in bicelles, and we achieved average resolution enhancements of 21–47% and individual peak enhancements as large as ca. 450%. New peaks were observed over the conventional NOESY experiment in both proteins as a result of these improvements, and the final structures generated from the calculated restraints matched published models. We discuss the impact of the experimental parameters, spin diffusion and the information content of the srNOESY lineshape.
The Lorieau Group (2018)
 Picture of Justin L Lorieau, Zoe Petros, Charles DeLisle, Indrani Banerjee, Alec Malooley, H. Bhagya Mendis and Medine Ayhan
Picture of Justin L Lorieau, Zoe Petros, Charles DeLisle, Indrani Banerjee, Alec Malooley, H. Bhagya Mendis and Medine Ayhan
Congratulations to Dr. Adrian Draney
 Congratulations to Adrian Draney on earning his PhD! Dr. Draney is the second
member of the Lorieau group to receive his PhD. He will join the
laboratory of Prof. Guido Pintacuda
at the Ecole normale supérieure de Lyon to
conduct solid-state NMR
experiments.
Congratulations to Adrian Draney on earning his PhD! Dr. Draney is the second
member of the Lorieau group to receive his PhD. He will join the
laboratory of Prof. Guido Pintacuda
at the Ecole normale supérieure de Lyon to
conduct solid-state NMR
experiments.
New Members Zoe Petros and Alec Malooley

 Welcome to Zoe
Petros and Alec Malooley, the newest graduate student members of the
Lorieau group. Zoe joins us from the University of Illinois,
Urbana-Champaign and Alec comes from the University of Illinois,
Chicago. They will be working on membrane protein structures by NMR.
Welcome to Zoe
Petros and Alec Malooley, the newest graduate student members of the
Lorieau group. Zoe joins us from the University of Illinois,
Urbana-Champaign and Alec comes from the University of Illinois,
Chicago. They will be working on membrane protein structures by NMR.
Congratulations to Dr. Sean Smrt
 Congratulations to Sean Smrt on earning his PhD! Dr. Smrt is the first
member of the Lorieau group to receive his PhD. He will join the
laboratory of Prof. Tim
Cross at the
National High Magnetic Field Laboratory at Florida State University to
conduct solid-state NMR
experiments on the membrane proteins of Mycobacterium tuberculosis.
Congratulations to Sean Smrt on earning his PhD! Dr. Smrt is the first
member of the Lorieau group to receive his PhD. He will join the
laboratory of Prof. Tim
Cross at the
National High Magnetic Field Laboratory at Florida State University to
conduct solid-state NMR
experiments on the membrane proteins of Mycobacterium tuberculosis.